|
|
|
|
|
|
|
|
link to the bioinformatic course (internal link) 
Publications: - Levin S., Sela N., Erez T., Nestel D., Pettis J., Neumann P., Chejanovsky N. (2019). New viruses in the ectoparasite mite Varroa destructor infesting Apis mellifera and Apis cerana. Viruses 11(2). pii: E94
- Maoz I., De Rosso M., Kaplunov T., Dalla Vedova A., Sela N., Flamini R., Lewinsohn E., Lichter A. (2019). Metabolomic and transcriptomic changes underlying cold and anaerobic stresses after storage of table grapes. Scientific reports
- Luria N., Smith E., Sela N., Koren A., Lachman O., Dombrovsky A.(2019). Insights into a watermelon virome contribute to monitoring distribution of whitefly borne viruses. Phytobiome
- Levitzky N., Smith E., Bakelman H., Luria N., Mizrahi Y., Lachman O., Sela N., Levi O., Milrot E., Dombrovsky A. (2019) The bumblebee Bombus terrestris contributes to Tomato brown rugose fruit virus spread in tomatoes. PLoS ONE 14(1):e0210871
- Katsir L., Zhepu R., Piasezky A., Jiang j., Sela N., Freilich S., Bahar O. (2018). Genome analysis of haplotype D of Candidatus Liberibacter solanacearum. Frontiers in microbiology
- Eliash N., Thangarajan S., Goldenberg I., Sela N., Kupervaser M., Barlev J., Altman Y., Kamer Y., Zaidman I., Rafaeli A., Soroker V (2018). Varroa chemosensory proteins: Some conserved across Arthropoda but others are Arachnid specific. Insect molecular biology
- Yakir E., Fei Z., Sela N., Xu Y., Singh V., Dagar A., Raj J.J., Müller M., Munné-Bosch S., Giovannoni J., Vrebalov J., Friedman H (2018). MaMADS2 repression in banana fruits modifies hormone synthesis and signaling pathways prior to climacteric stage. BMC Plant biology.
- Dror B., Savidor A., Salam B., Sela N., Lampert Y., Teper-Bamnolker P., Daus A., Carmeli S., Saldinger-Sela S., Eshel D (2018). High Levels of CO2 Induce Pathogenicity of Lactic Acid Bacteria by Upregulating Dextran-Synthesis Genes. Applied Environmental Microbiology.
- Massart S., Adams I., De Jonghe K., Giampetruzzi A., Koloniuk I., Kominek P., Kreuze J., Kutnjak D., Lotos L., Maree H.J., Thibaut O., Pooggin M., Ruiz-García A.B., Safarova D., Schneeberger P. H. H., Sela N., Varallyay E., Vainio E, Verdin E., Westenberg M., Brostaux Y., Candresse T. (2018) Virus detection by high throughput sequencing of small RNAs: large scale performance testing of sequence analysis strategies. Phytopathology
- Tannous J., Kumar D., Sela N., Sionov E., Prusky D., Keller N.P. (2018) Fungal attack and host defense pathways unveiled in near avirulent interactions of Penicillium expansum creA mutants on apples. Molecular Plant Pathology
- Maoz I., Davidovitch-Rikanati R., Shlezinger D., Bar E., Gonda I., Levin E., Kaplunov T., Sela N., Lichter A., Lewinsohn E (2018). Concealed ester formation and amino acids derive volatile compounds metabolism in table grape (Vitis vinifera L.) berries. Plant Science 274:223-230.
- Piombo E., Sela N., Wisniewski M., Hoffmann M., Gullino M.L., Allard M.W., Levin E., Spadaro D., Droby S.(2018) Genome draft of two Metschnikowia fructicola isolates - an important biocontrol agent delaying postharvest fruits decay. Frontiers in Microbiology. doi: 10.3389/fmicb.2018.00593
- Teixeira M.A., Sela N., Atamian H.S., Bao E., Chaudhary R., MacWilliams J., Mantelin S., Girke T., Kaloshian I. (2018). Sequence analysis of the potato aphid Macrosiphum euphorbiae transcriptome identified two new viruses. Accepted PLoS ONE.
- Luria N., Smith E., Sela N., Lachman O., Dombrovsky A. (2108). A local strain of Paprika mild mottle virus breaks L3 resistance in peppers and accelerates Tomato brown rugose fruit virus infection of Tm-22 resistant tomatoes. Virus Genes (2018): 1-10.
- Levin S., Galbraith D., Sela N., Erez T., Grozinger C., Chejanovsky N. (2017). Presence of Apis rhabdovirus-1 in populations of pollinators and their parasites from two continents. Front. Microbiol. doi: 10.3389/fmicb.2017.02482
- Katsir L., Zhepu R., Piasezky A., Jiang J., Sela N., Freilich S., Bahar O. (2017) The Genome Sequence of "Candidatus Carsonella ruddii" Strain BT, From the Psyllid Bactericera trigonica. Genome Announcement. In Press.
- Eliash N., Singh N.K., Raj S., Sela N., Leshkowitz D., Kamer Y., Zaidman I., Rafaeli A., Soroker V. (2017) Transcriptomic analysis of phoretic and reproductive olfactory organs of Varroa mite: Focus on chemosensory related genes. Scientific Reports 7: 13091
- Avrahami-Moyal L., Tam Y., Brumin M., Prakash S., Leibman D., Pearlsman M., Bornstein M., Sela N., Zeidan M., Dar Z., Zig U., Gal-On A., Gaba V. (2017) Detection of Potato virus Y in industrial quantities of seed potatoes by TaqMan Real Time PCR. Phytoparasitica DOI 10.1007/s12600-017-0612-
- Ofaim S., Ofek-Lalzar M., Sela N., Jiandong J., Kashi Y., Minz D., Frielich S (2017). Community-level metabolic-network analysis of functional division within microbial communities of plant roots. Front. Microbiol. 8:1606.
- Barad S., Sela N., Dubey A.K., Kumar D., Luria N., Ment D., Cohen S., Schaffer A.A., Prusky D. (2017). Differential gene expression in tomato fruit and Colletotrichum gloeosporioides during colonization of the RNAi-SlPH tomato line with reduced fruit acidity and higher pH. BMC Genomics (2017) 18:579
- Lampert Y., Dror B., Sela N., Teper-Bamnolker P., Zutahy Y., Sela (Saldinger) S., Eshel D. (2017). Emergence of Leuconostoc mesenteroides as a causative agent of oozing in carrots stored under non-ventilated conditions. Microbial Biotechnology DOI: 10.1111/1751-7915.12753
- Luria N.*, Smith E.*, Reingold V., Bekelman I., Lapidot M., Levin I., Elad N., Tam Y., Sela N., Abu-Ras A., Ezra N., Haberman A., Yitzhak L., Lachman O., Dombrovsky A (2017). A new Tobamovirus in Israel infects tomato plants harboring Tm-22 resistance genes. PLoS ONE 12(1): e0170429.
- Khankhum S., Sela N., Osorno J.M., Valverde R.A. (2016) RNAseq analysis of endornavirus-infected vs. endornavirus-free common bean (Phaseolus vulgaris) cv Black Turtle Soup. Front. Microbiol. 7:1905.doi: 10.3389/fmicb.2016.01905.
- Levin S., Sela N., Chejanovsky N. (2016). Two novel viruses associated with the Apis mellifera pathogenic mite Varroa destructor. Scientific Reports 6:37710 DOI: 10.1038/srep37710
- Trainin T., Zohar M., Shimoni-Shor E., Doron-Faigenboim A., Bar Ya'akov I., Hatib K.,Sela N., Holland D., Isaacson T (2016). A unique haplotype found in apple accessions exhibiting early bud-break could serve as a marker for breeding apples with low chilling requirements. Mol Breeding (2016) 36: 158. doi:10.1007/s11032-016-0575-7.
- Sivankalyani V., Sela N., Feygenberg O., Zemach H., Maurer D., Alkan N. (2016). Transcriptome Dynamics in Mango Fruit Peel Reveals Mechanisms of Chilling Stress. Front. Plant Sci. 7:1579. doi: 10.3389/fpls.2016.01579.
- Sela N., Luria N., Yaari M., Prusky D., Dombrovsky A. (2016). Genome Sequence of a Potential New Benyvirus, Isolated from Mango RNA-seq Data. Genome Announcements 4 (6):e01250-16.
- Buskila Y., Sela N., Teper-Bamnolker P., Tal I, Shani E., Weinstain R., Gaba V., Tam Y., Lers A., Eshel D. (2016). Stronger sink demand for gibberellin supports dominance of the apical bud in etiolated growth. Journal of Experimental Botany. August 31, 2016 doi:10.1093/jxb/erw315
- Barad S.*, Sela N.*, Verma D., Dubey A., Glam N., Sherman A., Prusky D. (2016) Fungal and host transcriptome analysis of pH regulated genes during colonization of Penicillium expansum on apple fruits. BMC Genomics 17:330 DOI: 10.1186/s12864-016-2665-7
- Luria N, Reingold V, Lachman O, Sela N, Dombrovsky A. (2016) Extended phylogenetic analysis of a new Israeli isolate of Brevicoryne brassicae virus (BrBV-IL) suggests taxonomic revision of the genus Iflavirus. virology journal 13:50. DOI: 10.1186/s12985-016-0500-z.
- Reingold V., Lachman O., Sela N., Luria N., Dombrovsky A. (2016) Watermelon fruit rot disease in Israel is caused by a distinct Squash vein yellowing virus (SqVYV) strain. Plant Disease. http://dx.doi.org/10.1094/PDIS-09-15-1040-RE.
- Teixeira M.,Sela N.,Ng J.,Casteel C.L.,Peng H.C.,Bekal S., Girke T.,Ghanim M.,Kaloshian I (2016). A novel virus from Macrosiphum euphorbiae with similarities to members of the family Flaviviridae. Journal of General Virology. 27 January, 2016 doi: 10.1099/jgv.0.000414
- Julca I., Droby S., Sela N., Marcet-Houben M., Gabaldón T. (2015) Contrasting genomic diversity in two closely related postharvest pathogens: Penicillium digitatum and Penicillium expansum. Genome Biol Evol Dec 14, 2015 doi:10.1093/gbe/evv252.
- Kaplan E, Sela N, Doron-faigenboim A, Navon-Venezia S, Jurkevitch E, Cytryn E. (2015) Genomic and functional characterization of qnr-encoding plasmids from municipal wastewater biosolid Klebsiella pneumoniae isolates. Front. Microbiol. 6:1354. doi: 10.3389/fmicb.2015.01354
- Ostrov I, Sela N, Freed M, Khateb N, Kott-Gutkowski M, Inbar D, Shemesh M. (2015) Draft genome sequence of Bacillus licheniformis S127 isolated from sheep udder clinical infection.Genome Announc 3(5):e00971-15. doi:10.1128/genomeA.00971-15.
- Iberkleid, I.,Sela N. and Brown-Miyara S. (2015). Meloidogyne javanica fatty acid- and retinol-binding protein (Mj-FAR-1) regulates expression of lipid-, cell wall-, stress- and phenylpropanoid-related genes during nematode infection of tomato. BMC Genomics 2015 16:272
- Levin, E., Sela N., Raphael G., Feygenberg O., Droby S., Wisniewski, M. (2014) Molecular Interactions between the Biocontrol Agent Metschnikowia fructicola and Citrus Fruit Tissue and Penicillium digitatum. Acta Hort. (ISHS) 1053:37-49.
- Ballester A.R., Marcet-Houben M., Levin E., Sela N., Selma-Lázaro C., Carmona L., Wisniewski M., Droby S., González-Candelas L., Gabaldón T. (2014). Genome, transcriptome, and functional analyses of Penicillium expansum provide new insights into secondary metabolism and pathogenicity. Molecular Plant-Microbe Interactions 2014 Oct 22.
- Lapidot M., Gelbart D., Gal-On A., Sela N., Anfoka G., Haj Ahmed F., Abou-Jawada Y., Sobh H., Mazyad H., Aboul-Ata A.E., El-Attar A.K., Ali-Shtayeh M.S., Jamous R.M., Polston J.E., Duffy S.,(2014) Frequent migration of introduced cucurbit-infecting begomoviruses among Middle Eastern countries. Virology journal 2014, 11:181.
- Luria N.*, Sela N.*, Yaari M., Feygenberg O., Kobiler I., Lers A., Prusky D. (2014) De-novo transcriptome assembly of mango fruit peel reveals mechanisms of mango response to hot water treatment. BMC Genomics 2014. Nov 5; 15:957 doi:10.1186/1471-2164-15-957 *equal contribution.
- Ofek-lalzar M., Sela N., Voronov-Goldman M., Green S.J., Hadar Y., Minz D., (2014) Niche and host-associated functional signatures of the root surface microbiome. Nature Communications 2014 Sep 18;5:4950. doi: 10.1038/ncomms5950
- Kolton M., Sela N., Elad Y., Cytryn E., (2013) Comparative genomic analysis indicates that niche adaptation of terrestrial Flavobacteria is strongly linked to plant glycan metabolism. (2013) PLoS ONE 8(9): e76704.
- Shemesh M., Pasvolsky R., Sela N., Green S.J., Zakin V. (2013) Draft genome sequence of Alicyclobacillus acidoterrestris strain ATCC 49025. Genome Announcement 2013 Sep 5;1(5).
- Rosenwasser S., Fluhr R., Joshi J.R., Leviatan N., Sela N., Hetzroni, A., Friedman H., (2013) ROSMETER: A bioinformatic tool for the identification ROS type and origin. Plant Physiol. August 2013 pp.113.218206.
- Dombrovsky A., Glanz E., Lachman O., Sela N., Doron-Faigenboim A., Antignus Y., (2013) The complete genome sequence of PYLCV and its implications on the understanding of evolution dynamics in the genus Polerovirus. (2013) PLoS ONE 2013 8(7): e70722.
- Sela N., Lachman O., Reingold V., Dombrovsky A., (2013) A new cryptic virus belonging to the family Partitiviridae was found in watermelon co-infected with Melon necrotic spot virus. 2013 Virus Genes June 18. DOI: 10.1007/s11262-013-0937-8.
- Droby, S., Hershkovitz, V., Sela, N., Raphael, G., Kessler, C., Feygenberg, O. and Wisniewski, M. (2013). Biochemical and transcriptomic analysis of host responses to yeast biocontrol agents of postharvest pathogens. Acta Hort. (ISHS) 1012:671-680
- Hershkovitz V.*, Sela N.*, Taha- Salaime L., Liu J., Rafael G., BenDayan C., Aly R., Wisniewski M., Droby S. (2013) De-novo assembly and Characterization of the Transcriptome of Metschnikowia fructicola reveals differences in gene expression following interaction with Penicillium digitatum and grapefruit peel. BMC Genomics 2013, 14:168 *equal contribution
- Kolton M., Green S.J., Meller-Harel Y., Sela N., Elad Y., Cytryn E.,(2012) Draft Genome Sequence of Flavobacterium sp. Strain F52, Isolated from the Rhizosphere of Red Pepper (Capsicum annuum L). 2012 Journal of Bacteriology 194 (19), 5462-3.
- Sela N., Luria N, Dombrovsky A., (2012) Genome assembly of Bell pepper endornavirus from small RNA. 2012 Journal of virology Vol. 86 doi:10.1128/JVI.00983-12
- Blum S., Sela N., Heller H.D., Sela S., Leitner G., (2012) Genome analysis of bovine mastitis associated Escherichia coli O32:H37 strain P4. J. Bacteriol. 2012 vol. 194 no. 14 3732
- Madrigal P., Sela N., Lin S.M., (2011) PLoS Computational Biology Conference Postcards from ISMB/ECCB 2011. PLoS Computational Biology 2011 Nov;7(11):e1002259
- Sela N., Mersch B., Hotz-Wagenblatt A., Ast G., (2010) New Characteristics of transposable element exonization within human and mouse. PLoS ONE 2010; 5(6):e10907.
- Sela N., Kim E., Ast G., (2010) The role of transposable elements in the evolution of non-mammalian vertebrates and invertebrates. Genome Biology 2010; 11(6):R59.
- Sela N., Stern A., Makalowski W., Pupko T., Ast G., (2008) Transduplication resulted in the incorporation of two protein-coding sequences into the Turmoil-1 transposable element of C. elegans. Biology Direct 2008; 3:41.
- Lev-Maor G.*, Ram O.*, Kim E.*, Sela N.*, Goren A., Levanon E.Y., Ast G., (2008) Intronic Alus influence alternative splicing. PLoS Genetics, 2008; 4(9):e1000204 *equal contribution
- Levy A.*, Sela N.*, Ast G.,(2008) TranspoGene and MicroTranspoGene: Transposed elements influence on the transcriptome of seven vertebrates and invertebrates. Nucleic Acid Research, 2008; 36 (Database issue): D47-D52 *equal contribution
- Amit M., Sela N., Keren H., Melamed Z., Muler I., Shomron N., Izraeli S., Ast G., (2007) Biased exonization of transposable elements in duplicated genes: a lesson from TIF-IA gene; BMC Molecular Biology, 2007; 8:109.
- Mersch B., Sela N., Ast G., Suhai S., Hotz-Wagenblatt A.,(2007) SERpredict: Detection of tissue- or tumor-specific isoforms generated through exonization of transposable elements. BMC genetics, 2007; 8: 78.
- Lev-Maor G.*, Goren A.*, Sela N.*, Kim E., Keren H., Doron-Faigenboim A., Leibman-Barak S., Pupko T., Ast G. (2007) The "Alternative" choice of constitutive exons through evolution. PLoS Genetics, 2007; 3(11): e203 *equal contribution
- Sela N., Mersch B., Gal-Mark N., Lev-Maor G., Hotz-Wagenblatt A., Ast G.,(2007) Comparative analysis of transposed elements' insertion within human and mouse genomes reveals Alu's unique role in shaping the human transcriptome; Genome Biology, 2007; 8(6):R127.
- Sela N., Degani H., Frydman L.,(2004) Ultrafast 2D NMR using sinusoidal gradients: principles and ex-vivo brain investigations; Magnetic Resonance in Medicine, 2004 Oct; 52(4):893-7.
- Dadiani M., Margalit R., Sela N., Degani H.,(2004) High Resolution Magnetic Resonance imaging of Disparities in the Transcapillary Transfer Rates in Orthotopically Inoculated Invasive Breast Tumors; Cancer Research, , 2004 May 1; 64 (9): 3155-61.
|
|
|
|
|
|
|
 |
|
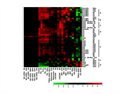
ROSMETER: A new bioinformatic tool |
| Oxidative stress in different organelles lead to a different transcriptomic imprint. We developed a new bioinformatic tool, ROSMETER, that is able to identify the ROS-related transcriptome signature related to stresses in different organelles. |
| 0 Comments |
|
|
|
|
| See all chapters |
|
|
| Updated on: 07/02/24 16:54 |
|
|
|
|
|
|
|